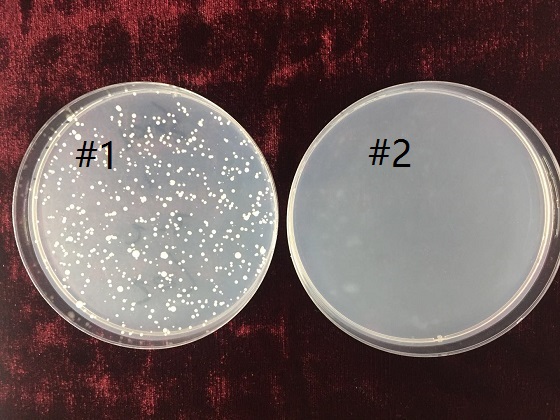
A yeast genetic system for selecting small molecule inhibitors of protein–protein interactions in nanodroplets | PNAS

Generation of the barcode-taggedS. pombeinsertion mutant library. The... | Download Scientific Diagram
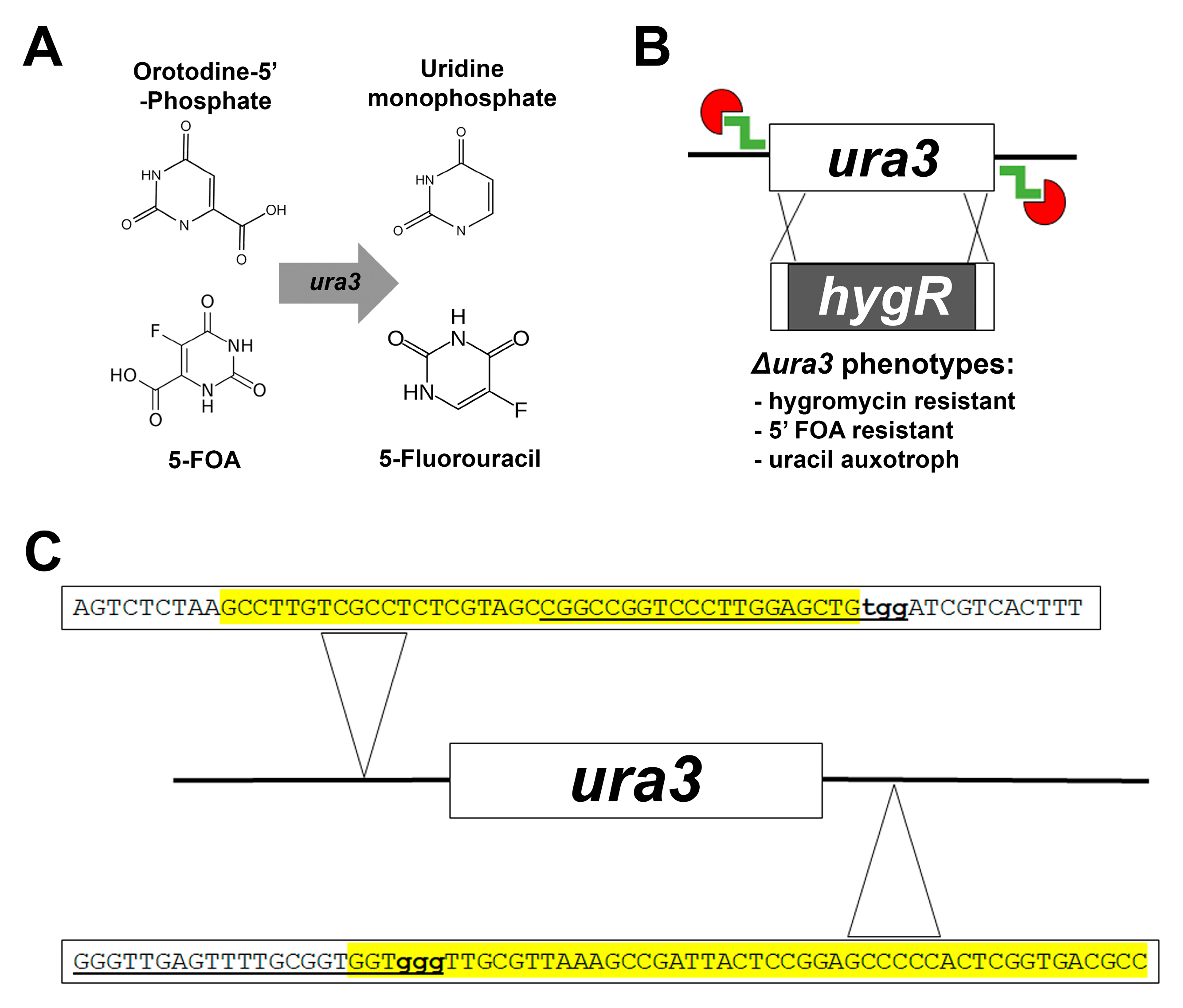
Microorganisms | Free Full-Text | CRISPR/Cas9-Mediated Gene Replacement in the Fungal Keratitis Pathogen Fusarium solani var. petroliphilum
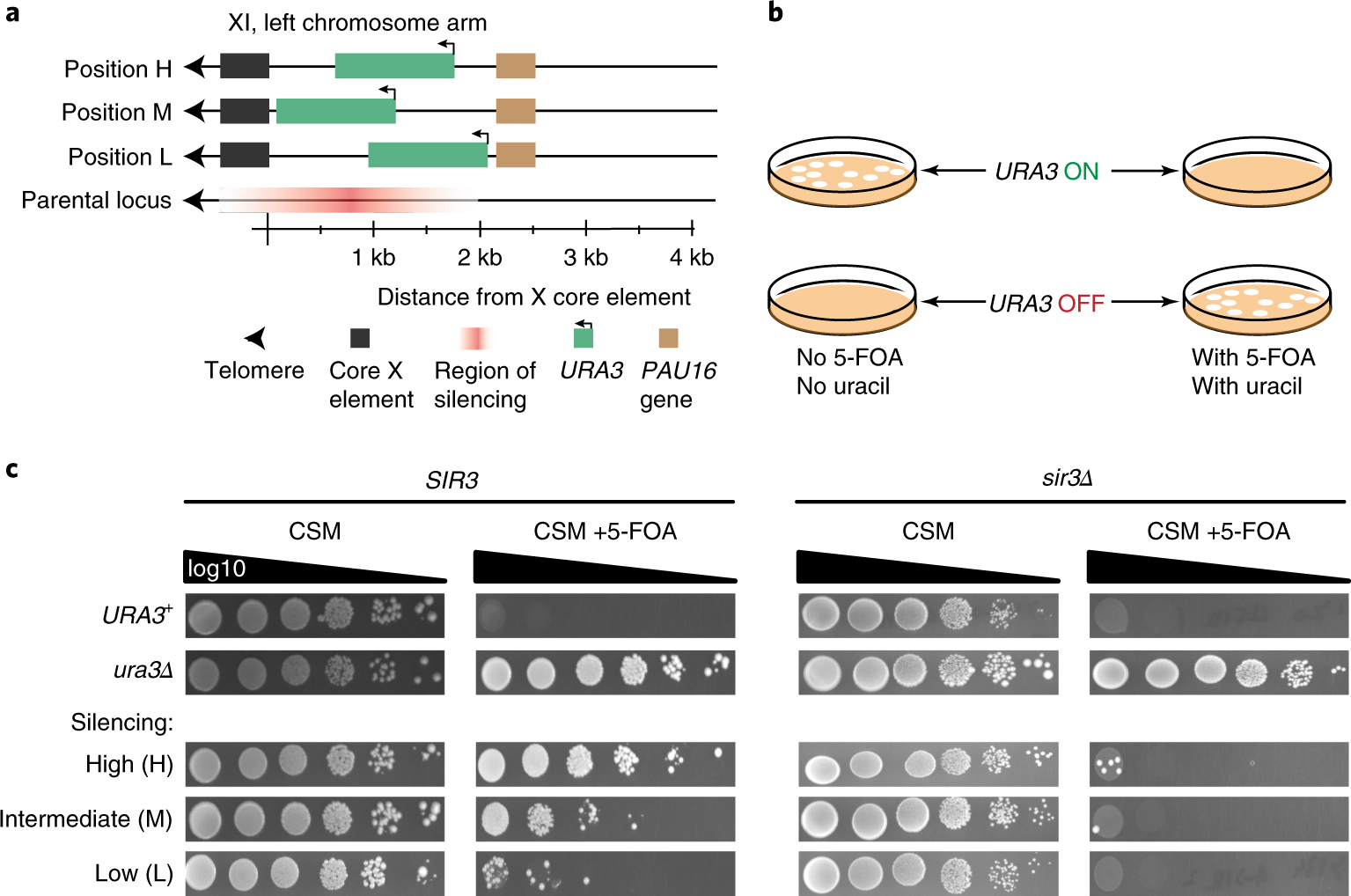
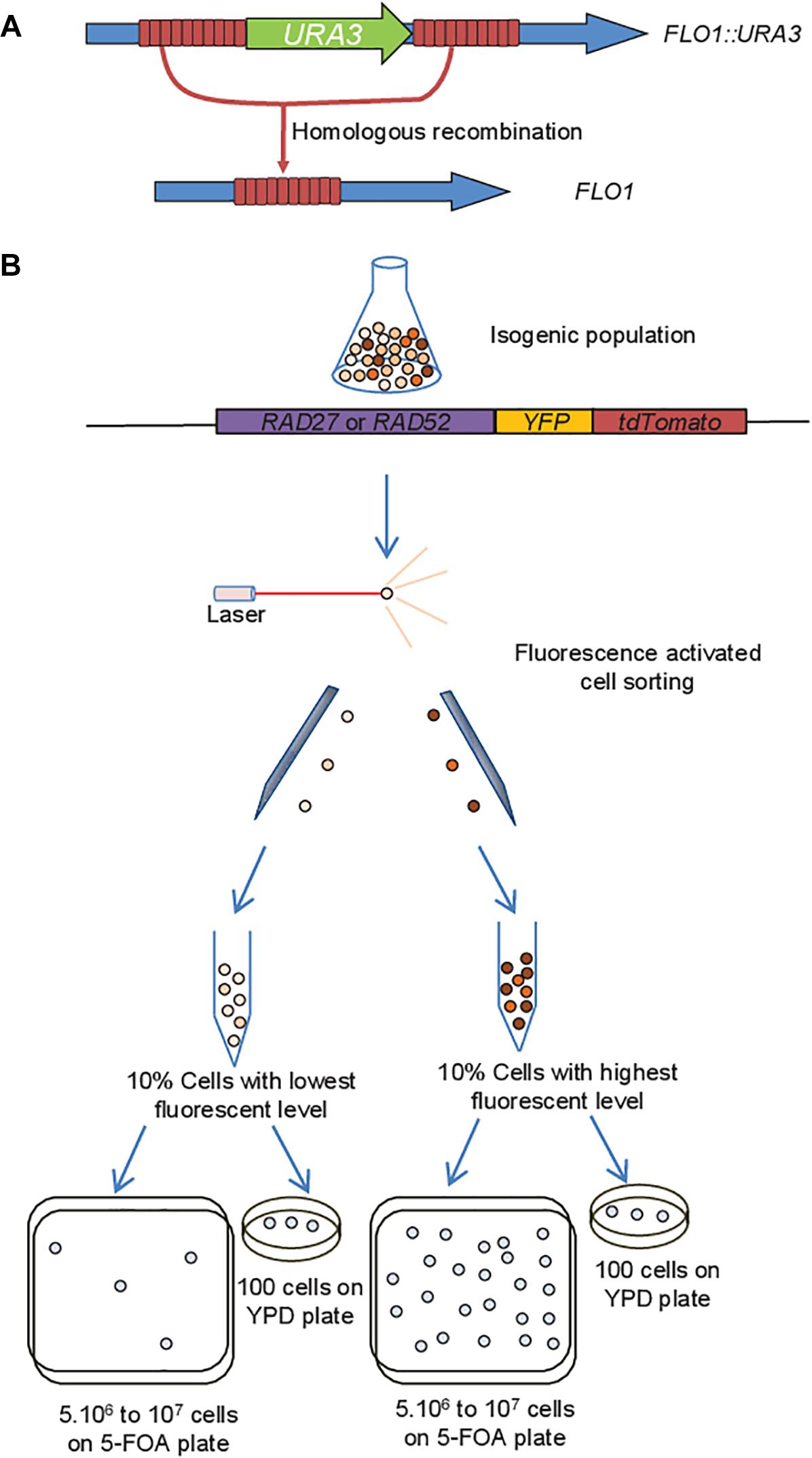
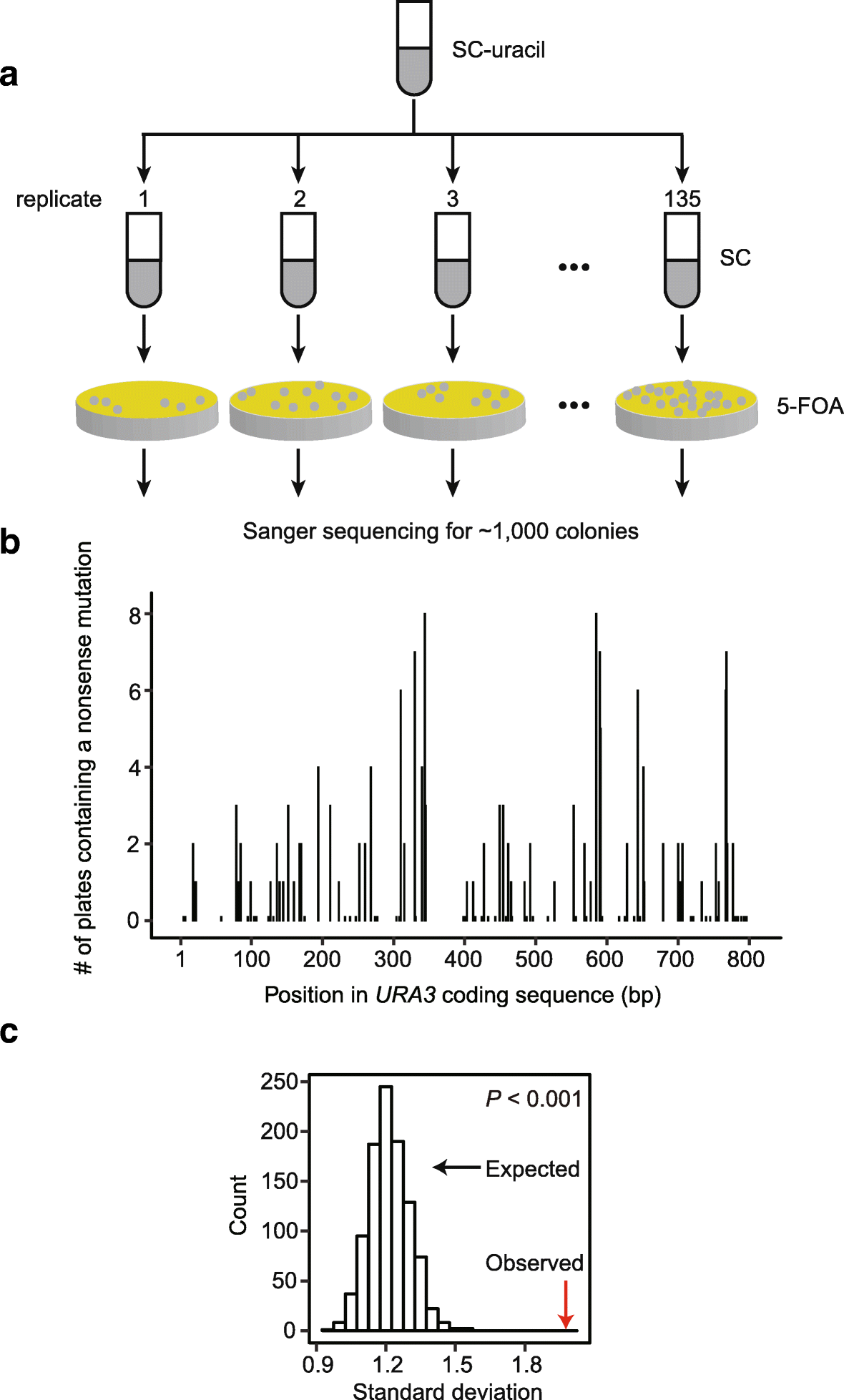
Epigenetic gene silencing alters the mechanisms and rate of evolutionary adaptation | Nature Ecology & Evolution
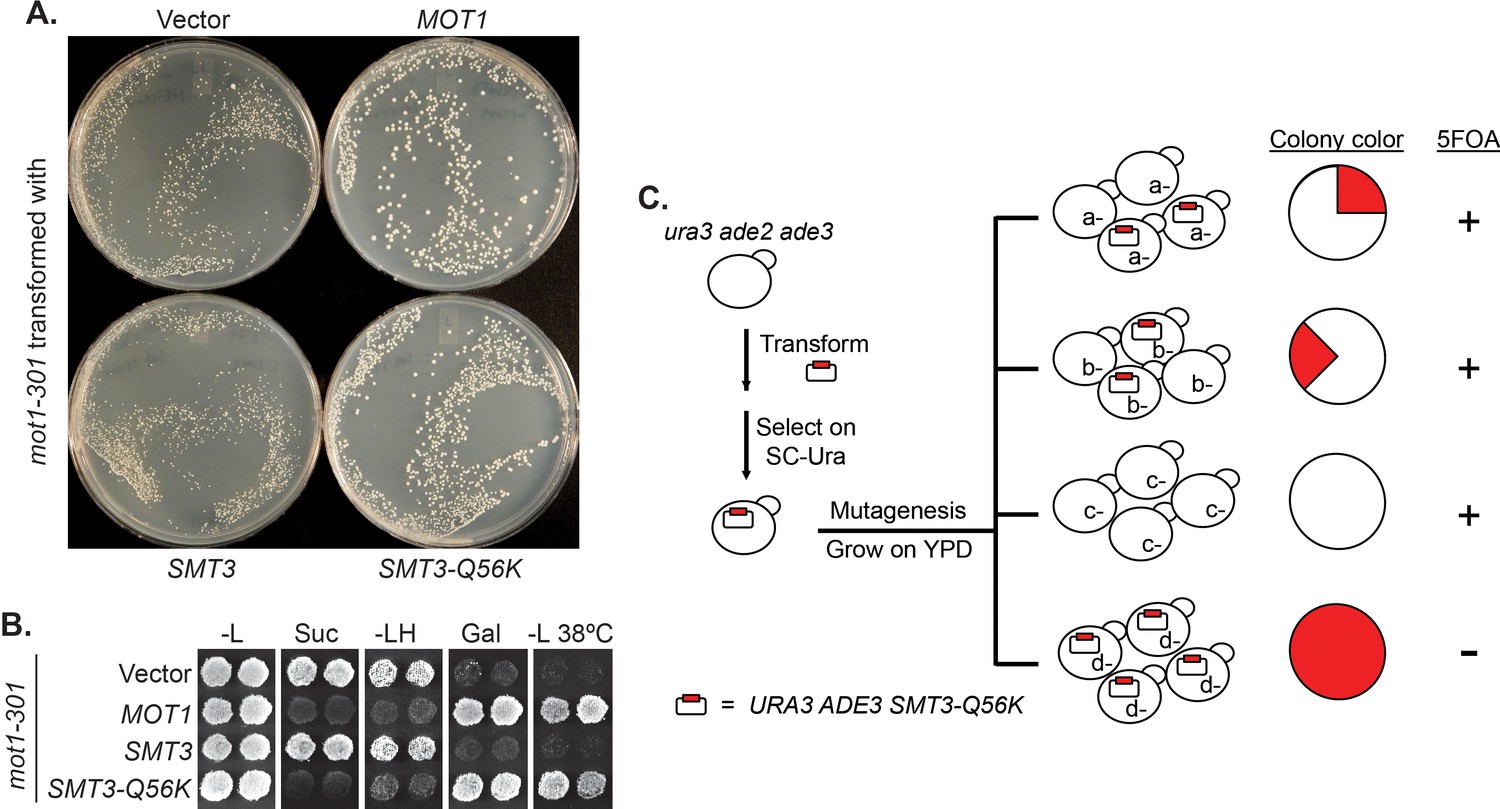
Figures and data in Defective RNA polymerase III is negatively regulated by the SUMO-Ubiquitin-Cdc48 pathway | eLife

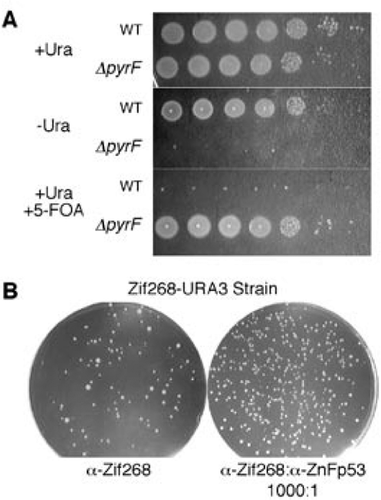
Phenotyping transformants for uracil auxotrophy and 5-FOA resistance.... | Download Scientific Diagram

Development of a pyrG Mutant of Aspergillus oryzae Strain S1 as a Host for the Production of Heterologous Proteins